Overview
Croizat is a free, user-friendly, cross-platform desktop software package which biologists can use to integrate and analyze spatial data on species or other taxa and to explore geographical patterns in diversity under a panbiogeographic and graph-theoretic approach.
Panbiogeography provides a method for analyzing the geographic structure of distributions in order to generate predictions about the evolution of species and other taxa in space and time. As of today, there is no standard, general-purpose software for the analysis of distributional data under a panbiogeographic approach. Croizat aims to solve that. It is based on the same database/analytical tools/map graphics model of many Geographic Information Systems (GIS). It does not have all the features and functions of a GIS, but this makes it much easier to use. Yet unlike GIS's, rather than concentrating on database and graphics flexibility, Croizat is designed to perform specialized biological analyses, many of which are not readily available from GIS's.
Features 
- An easy-to-use, interactive Graphical User Interface (GUI), with pulldown menus, dialog boxes, and other standard GUI controls, with almost identical look-and-feel on GNU/Linux, Microsoft Windows, and Apple Macintosh personal computers.
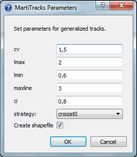
- Track analysis, with fast computing of individual tracks as minimum-spanning trees from a distance matrix among stations, as well as obtanining sets of compatible tracks (generalized tracks), using the MartiTracks program which is automatically run under the hood.
- Data import in a variety of formats, including Excel and Calc spreadsheets, dBase files, tab- and comma-delimited text generated by all the major online biodiversity databases (GBIF, iDigBio, OBIS, VertNet), as well as standard GIS shapefiles and KML files used by Google Earth.
- Output of shapefiles and KML files readable for map display with any Geographic Information System (DIVA-GIS, QuantumGIS, ArcGIS).
- Locality records are displayed as symbols (squares, circles, crosses, etc) on the map, with different symbols for each species.
- Optional overlay of satellite imagery on maps, using NASA "Blue Marble" images.
- Optional display of rivers and country boundaries.
- Zoom in and out on areas of interest.
- Point, line and fill colours can be set.
- Size, colour and type of symbols can be set.
- Publication-quality maps, that can be saved as graphics files, or copied to the clipboard and pasted into other applications.
Additional features (as direct seach and data retrieval from online biodiversity databases and grid analysis for locating of main massings) will be included in future releases.
Croizat is written in Python, an interpreted, interactive, object-oriented programming language, coupled with the portable, multi-platform Qt interface management library, and other free external libraries also written in Python and C/C++ (Scipy, NumPy, Matplotlib and its Basemap module). Therefore, the program is platform-independent, and runs without modifications on any PC compatible with the x86 architecture, under GNU/Linux, Mac OS X, and MS-Windows.
Screenshots 
Click on the images to enlarge.
 |
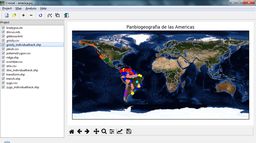 |
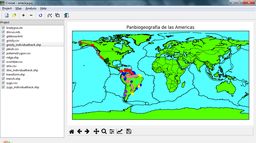
| Croizat main screen (default view) | Croizat main screen (using NASA "Blue Marble" image as map background) |
| Croizat/MartiTracks parameters dialog | Distribution of Zygodontomys rodents in northern South America represented as a track graph computed by Croizat (with satellite image background) |
Download & Installation 
Source code and binary installaton packages are available from SourceForge. There is no need to download external files and perform complicated system configurations; everything needed is already included in the installation package. Notice: the binary installation package is available only for MS-Windows at this time.
To install the software, just download and execute the binary installation package. The installer places a double-clickable executable in the Croizat folder inside your Program directory, together with the example files and the documentation. To start the program, simply double-click on the executable.
The software is distributed as free software, under the GNU General Public License (GPL).
Lite Version 
A "lite" version of the software (Croizat Reloaded), with limited functionality, is also available. It focuses on analysing data and generating output files, with all actual map display being done by means of a Geographic Information System (DIVA-GIS, Quantum GIS, ArcGIS, etc.).
Documentation & Support 
The user's guide for Croizat is included in the installation package and also available here.
It is recommendable to join the Panbiog-L discussion list for discussions and exchanging questions about the software, as well as to get information about updates and bugs.
Citation 
The correct citation suggested for Croizat in technical and scientific publications is as follows:
Applications 
Scientific works which have used Croizat:
- Santos-Silva, E. N., Brandorff, G.-O. and Cavalcanti, M. J. 2019. Distribution of European and African species of genus Diaptomus (Copepoda: Calanoida: Diaptomidae): a track analysis. Nauplius 26: e2018028.
- Cavalcanti, M. J., Lopes. P. R. D. and Sanos-Silva, E. N. 2019. Panbiogeographic analysis of the distribution patterns of the freshwater species of Sciaenidae (Actinopterygii: Perciformes) in the Amazon Basin. Tropical Diversity 1(1): 5-11.
- Cruz da Silva, E. L. 2013. Description of the male of Architis catuaba Santos & Nogueira, 2008 and new records of Architis capricorna Carico, 1981 (Araneae: Pisauridae). Zootaxa 3635(4): 485–490.
- Moreira, G.R.P., Ferrari, A., Mondin, C. and Cervi, A. 2011. Panbiogeographical analysis of passion vines at their southern limit of distribution in the Neotropics. Revista Brasileira de Biociências 9: 28-40.
- Previatelli, D. 2010. Phylogeny and biogeography of Neotropical Diaptominae. Unpublished PhD Thesis, Instituto Nacional de Pesquisas da Amazônia, Manaus. 204p.
In the News 
- Conservation International Press Release
- Distance Learning in Developing Countries Project Overview
- Happy News Notice
Twitter
Updates 
 Be notified of updates to Croizat
by following us on
Twitter.
Be notified of updates to Croizat
by following us on
Twitter.
Acknowledgements

Many people have contributed to the development of Croizat: Brad McFall, Claudio Quezada, Daniel Previatelli, Douglas Riff, Edinaldo Nelson dos Santos-Silva, Gervásio Carvalho, Ian Henderson, John Grehan, Jonathan Liria. José Maria Cardoso da Silva, Juan Morrone, Maria Lucia Lorini, Michael Heads, Nicholas Matzke, Renato Bernils, Ronaldo Alperin, Tania Escalante, and Valéria Gallo. Jeffrey Whitaker, Scott Sinclair, John Hunter (in memoriam), Ryan May, Stef Mientki, and Benjamin Root (members of the Matplotlib-users mailing list) provided assistance and technical expertise. Special thanks to Robin Craw for encouragment and moral support. The development of Croizat has been funded by grants from Conservation International Brazil (version 1.x) and Projeto Biotupé, Instituto Nacional de Pesquisas da Amazônia (version 2.x).
Contact 
Mauro J.
Cavalcanti
P.O. Box 18123
CEP 20720-970
Rio de Janeiro, RJ, BRAZIL
Copyright © 2003-2021
|
Last updated November 8, 2021 |
Sponsored by  |
Contribute with a donation and help to keep alive the development of Croizat. Donations can be made with credit card, using PayPal.